
University of Nebraska-Lincoln
729(Lab) 551A(Vice Chair Office) Hamilton Hall
Lincoln, NE 68588-0304
(402) 472-2998 (Lab)
(402) 472-3502 (Vice Chair Office)
ldu3@unl.edu
Education
Postdoc. University of California, Davis
Ph.D., The Royal Vet and Ag University, Denmark
M.S., The Chinese Academy of Sciences
B.S., Yunnan University, China
Research Interests
Biosynthetic mechanism and metabolic engineering of novel antifungal and anti-MRSA antibiotics from gliding Gram-negative bacteria, mycotoxins and other bioactive natural products from plant pathogenic fungi and endophytes
Current Research
- Molecular mechanism for the biosynthesis of new antibiotics, mycotoxins and other bioactive natural products
- Discovery of new antibiotics and other bioactive natural products from Lysobacter bacteria, endophytic fungi, and other underexplored sources
- Metabolic pathway engineering
Research in the Du group is at the interface of chemistry and biology. Specifically, we are studying the molecular mechanisms by which organisms (bacteria, fungi and plants) make structurally complex, biologically active natural products. Our goal is to use this knowledge to produce new products through genetic engineering that will be beneficial to humans. The studies involve tools and knowledge from biochemistry, molecular biology, genetics, and chemistry. Currently, we are working on three main projects.
1. Biosynthetic mechanism for fungal polyketides
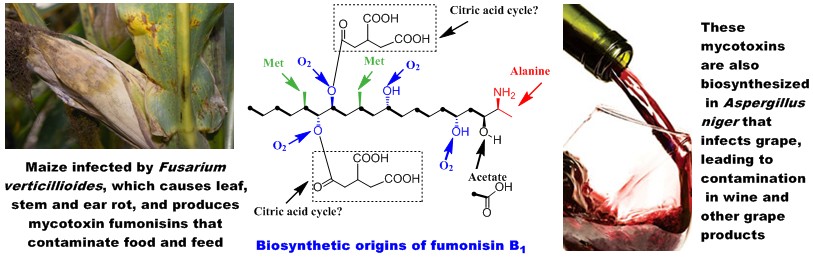
Polyketides are probably the most significant group of natural products in terms of their importance to human medicines. To date, most of the studies have focused on polyketides isolated from bacteria. Although fungi produce numerous polyketides, their biosynthetic mechanism is still not very clear. We have chosen a group of polyketides, including mycotoxin fumonisins, as a model system for our studies. Fumonisins are produced by the pathogenic fungus Fusarium verticillioides, which is a widespread contaminant of corn and maize-derived food and feed (Biopolymers 2010). The ingestion of fumonisin-contaminated corn causes fatal diseases in livestock and imposes cancer risk to humans. We have developed a genetic system that can specifically change the biosynthetic genes in filamentous fungi. Using this system, we have created mutants for the biosynthetic genes, determined biosynthetic intermediates in the mutants, and established a biosynthetic pathway. Through engineering the polyketide synthase gene, we successfully turned the mycotoxin-producing fungus into an antifungal metabolite-producer (JACS 2007). In addition, we have been using E. coli and baker's yeast as heterologous hosts to express and study the fungal genes (Biochem 2006). Our studies revealed an unprecedented PLP-dependent polyketide chain-releasing mechanism, in which a discrete 2-oxoamine synthase catalyzes a decarboxylative condensation between L-alanine and acyl-S-ACP (JACS 2009). The reaction results in the termination and offloading of the polyketide chain, as well as the introduction of a new carbon-carbon bond and an amino group to the chain. The mechanism is fundamentally different from the thioesterase/cyclase-catalyzed polyketide chain-releasing mechanism found in bacterial and other fungal polyketide biosyntheses (NPR 2010).
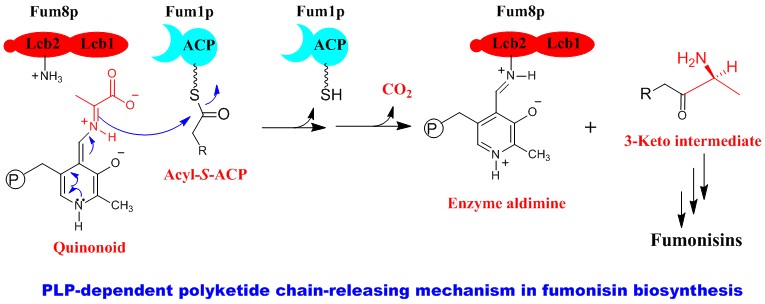
2. New antibiotics from a group of ubiquitous environmental bacteria
Lysobacter is a genus of Gram-negative bacteria that are ubiquitous in soil and aquatic environments. Several Lysobacter species are prolific producers of bioactive natural products and emerging as new biocontrol agents (NPR 2012). From the biocontrol agent Lysobacter enzymogenes, we have isolated a potent antifungal compound HSAF, which appears to have a novel mode of action (MBC 2006; RSC Adv 2016; BBA 2016) and possesses new structural features distinct from any existing fungicides and antifungal drugs (AAC 2007). We have identified the genes for HSAF biosynthesis and investigated the biosynthetic mechanism (JACS 2011; Biochem 2012; Angew Chem 2014; AMB 2016). These studies revealed a previously unrecognized biosynthetic mechanism for hybrid polyketide-peptide. In addition, we isolated a group of potent anti-MRSA cyclic lipodepsipeptides WAP-8294A and identified the WAP biosynthetic gene cluster from the bacterium (AAC 2011; RSC Adv 2015). These works establish the foundation for exploitation of Lysobacter species for new antibiotics (ACS Synth Biol 2013).More recently, we have looked into regulatory mechanisms for the biosynthesis (AMB 2015) and revealed a flavin-dependent aromatic N-oxide formation in phenazine antibiotics (Org Lett 2016) .
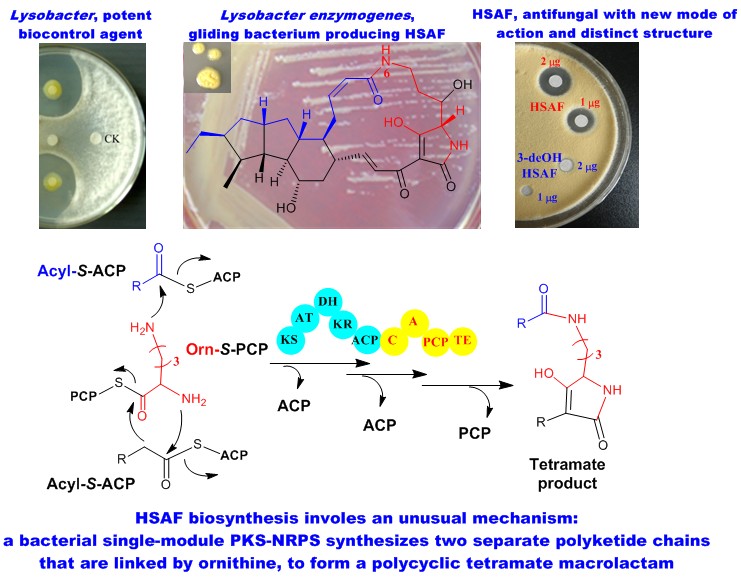
3. New Anticancer Natural Products from Plant Endophytic Fungi
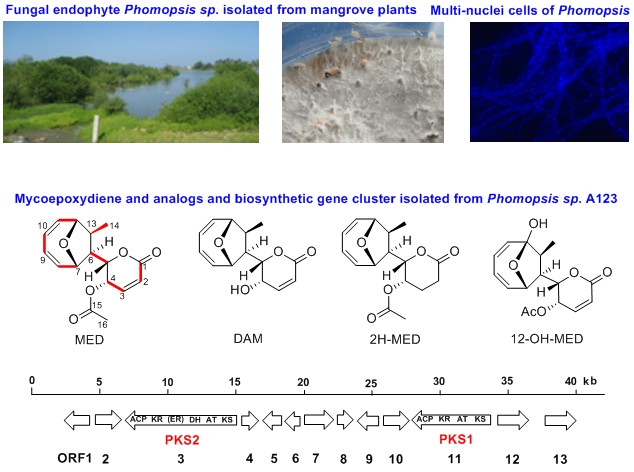
Plant endophytic microorganisms represent a largely unexploited resource for new bioactive natural products. It is estimated that each of the ~300,000 plant species on the earth has at least one endophyte. Collaborating with Prof. Yuemao Shen at the School of Life Science, Shandong University, we have isolated a family of structurally distinct and biologically active natural products from a new marine fungal strain, Phomopsis sp A123, which is an endophyte of the costal mangrove plants. These compounds, including mycoepoxydiene (MED) and deacetylmycoepoxydiene (DAM), exhibit a variety of biological properties including anticancer, anti-inflammatory and antimicrobial activities (AMB 2016) . Most interestingly, the compounds possess a rare structural feature, a cyclooctadiene with an oxygen bridge. To investigate the molecular mechanism for the biosynthesis of these compounds, we have cloned the biosynthetic genes from the endophytic fungus and used RNAi to verify the genes.
For more information, please visit the Du Research Group Homepage.
Selected Publications
(24) Yue H, Miller A, Khetrapal V, Jayasekera V, Wright S, and Du L. 2022. Biosynthesis, regulation, and engineering of natural products from Lysobacter. Natural Product Reports in press, DOI: 10.1039/D1NP00063B.
(23) Yue H, Jiang J, Taylor A, Leite A, Dodds E, and Du L. 2021. Outer membrane vesicles-mediated co-delivery of the antifungal HSAF metabolites and lytic polysaccharide monooxygenase in the predatory Lysobacter enzymogenes. ACS Chemical Biology 16: 1079-1089. pdf
(22) Yu L, Du F, Chen X, Zheng Y, Morton M, Liu F, and Du L. 2020. Identification of the biosynthetic gene cluster for the anti-MRSA lysocins through gene cluster activation using strong promoters of housekeeping genes and production of new analogs in Lysobacter sp. 3655. ACS Synthetic Biology 9: 1989-1997. pdf
(21) Sang M, Wang H, Shen Y, Rodrigues de Almeida N, Conda-Sheridan M, Li S, Li Y, and Du L. 2019. Identification of an anti-MRSA cyclic lipodepsipeptide, WBP-29479A1, by genome mining of Lysobacter antibioticus. Organic Letters 21: 6432-6436. pdf.
(20) Li X, Wang H, Shen Y, Li Y, and Du L. 2019. OX4 is an NADPH-dependent dehydrogenase catalyzing an extended Michael addition reaction to form the six-membered ring in the antifungal HSAF. Biochemistry 58: 5245−5248. pdf
(19) Li Y, Wang H, Liu Y, Jiao Y, Li S, Shen Y, and Du L. 2018. Biosynthesis of the polycyclic system in the antifungal HSAF and analogues from Lysobacter enzymogenes. Angewandte Chemie International Edition 57: 6221-6225. pdf
(18) Jiang J, Guiza-Beltran D, Schacht A, Wright S, Zhang L, and Du L. 2018. Functional and structural analysis of phenazine O-methyltransferase LaPhzM from Lysobacter antibioticus OH13 and one-pot enzymatic synthesis of the antibiotic myxin. ACS Chemical Biology 13: 1003-1012. pdf.
(17) Yu L, Su W, Fey P, Liu F, and Du L. 2018. Yield improvement of the anti-MRSA antibiotics WAP-8294A by CRISPR/dCas9 combined with refactoring self-protection genes in Lysobacter enzymogenes OH11. ACS Synthetic Biology 7: 258−266. pdf.
(16) Li S, Wu X, L Zhang, Shen Y, and Du L. 2017. Activation of a cryptic gene cluster in Lysobacter enzymogenes revealed a module/domain portable mechanism of nonribosomal peptide synthetases in the biosynthesis of pyrrolopyrazines. Organic Letters 19: 5010−5013. pdf.
(15) Han Y, Wang Y, Yu Y, Chen H, Shen Y, and Du L. 2017. Indole-induced reversion of intrinsic multi-antibiotic resistance in Lysobacter enzymogenes. Applied and Environmental Microbiology 83:e00995-17. pdf.
(14) Zhao Y, Qian G, Ye Y, Wright S, Chen H, Shen Y, Liu F, and Du L. 2016. Heterocyclic aromatic N-oxidation in the biosynthesis of phenazine antibiotics from Lysobacter antibioticus. Organic Letters 18: 2495-2498. pdf
(13) Wang M, Zhang W, Xu W, Shen Y, and Du L. 2016. Optimization of genome shuffling for high yield production of the antitumor deacetylmycoepoxydiene in an endophytic fungus of mangrove plants. Applied Microbiology and Biotechnology 100: 7491–7498. pdf
(12) Han Y, Wang Y, Tombosa S, Wright S, Huffman J, Yuen G, Qian G, Liu F, Shen Y, and Du L. 2015. Identification of a small molecule signaling factor that regulates the biosynthesis of the antifungal polycyclic tetramate macrolactam HSAF in Lysobacter enzymogenes. Applied Microbiology and Biotechnology 99: 801-811. pdf
(11) Li Y, Chen H, Ding Y, Xie Y, Wang H, Cerny RL, Shen Y, and Du L. 2014. Iterative assembly of two separate polyketide chains by the same single-module bacterial polyketide synthase in the biosynthesis of HSAF. Angewandte Chemie International Edition 53: 7524-7530. pdf
(10) Wang Y, Qian G, Liu F, Li Y-Z, Shen Y, and Du L. 2013. Facile method for site-specific gene integration in Lysobacter enzymogenes for yield improvement of the anti-MRSA antibiotics WAP-8294A and the antifungal antibiotic HSAF. ACS Synthetic Biology 2: 670-678. pdf
(9) Xie Y, Wright S, Shen Y, and Du L. 2012. Bioactive natural products from Lysobacter. Natural Product Reports 29: 1277-1287. pdf
(8) Lou L, Chen H, Cerny RL, Li Y, Shen Y, and Du L. 2012. Unusual activities of the thioesterase domain for the biosynthesis of the polycyclic tetramate macrolactam HSAF in Lysobacter enzymogenes C3. Biochemistry 51, 4-6. pdf. (Highlighted on Biochemistry website, January, 2012).
(7) Zhang W, Li Y, Qian G, Wang Y, Chen H, Li Y-Z, Liu F, Shen Y, and Du L. 2011. Identification and characterization of the anti-Methicillin-Resistant Staphylococcus aureus WAP-8294A2 biosynthetic gene cluster from Lysobacter enzymogenes OH11. Antimicrobial Agents and Chemotherapy 55: 5581-5589. pdf.
(6) Lou L, Qian G, Xie Y, Hang J, Chen H, Zaleta-Rivera K, Li Y, Shen Y, Dussault PH, Liu F, and Du L. 2011. Biosynthesis of HSAF, a tetramic acid-containing macrolactam from Lysobacter enzymogenes. Journal of the American Chemical Society 133: 643-645. pdf.
(5) Du L and Lou L. 2010. PKSs and NRPSs release mechanisms. Natural Product Reports 27: 255-278. pdf.
(4) Gerber R, Lou L, and Du L. 2009. A PLP-dependent polyketide chain releasing mechanism in the biosynthesis of mycotoxin fumonisins in Fusarium verticillioides. Journal of the American Chemical Society 131: 3148-3149. pdf. (Highlighted on JACS website, March, 2009).
(3) Zhu X, Yu F, Li X-C, and Du L. 2007. Production of dihydroisocoumarins in Fusarium verticillioides by swapping the ketosynthase domain of the fungal iterative modular polyketide synthase Fum1p with that of lovastatin diketide synthase. Journal of the American Chemical Society 129: 36-37. pdf.
(2) Yu F, Zaleta-Rivera K, Zhu X, Huffman J, Millet J, Harris SD, Yuen G, Li X, and Du L. 2007. Structure and biosynthesis of HSAF, a broad spectrum antimycotic with a novel mode of action. Antimicrobial Agents and Chemotherapy 51: 64-72. pdf. Cited by Faculty of 1000 Biology (Evaluated by D. Newman, NIH, March 8, 2007) pdf.
(1) Zaleta-Rivera K, Xu C, Yu F, Butchko RAE, Proctor RH, Lara MEH, Raza A, Dussault PH, and Du L. 2006. A bidomain nonribosomal peptide synthetase encoded by FUM14 catalyzes the formation of tricarballylic esters in the biosynthesis of fumonisins. Biochemistry 45: 2561-2569. pdf