
University of Nebraska-Lincoln
431 Hamilton Hall
Lincoln, NE 68588-0304
(402) 472-5232
hli4@unl.edu
Education
Postdoctoral Associate, Iowa State University
Ph.D., University of Iowa
B.Sc., Lanzhou University
Teaching and Research Interests
Quantum chemistry and QM/MM methods
Current Research
Structural Biology. Large-scale second-order perturbation theory (MP2) calculations are used to determine accurate active site structures in biological molecules. Such structural and energetics information can greatly help us understand the active site chemistry.
Metalloprotein Redox Chemistry. The reduction potentials (E0) of transition metal ions (such as Fe3+ and Cu2+) in metalloproteins are calculated using quantum methods, and the protein regulations on the E0 are analyzed.
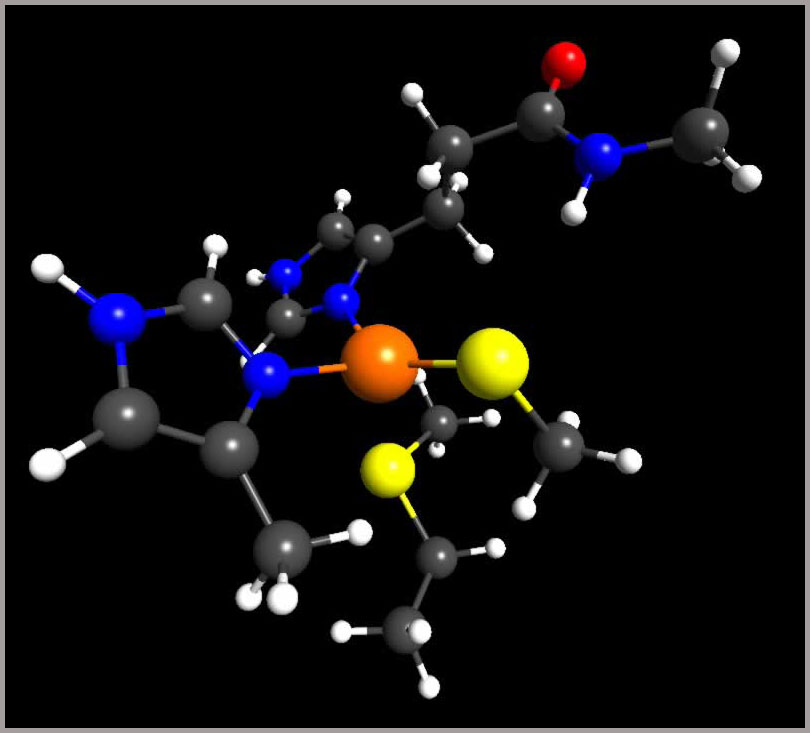
Histidine pKa. Among the ionizable groups in proteins, histidine exhibit the most complicated pKa chemistry, especially those bound to metal ions, or involved in catalytic active sites. We perform accurate quantum chemical calculations to analyze how hydrogen bonding and solvation effects tune histidine pKa.
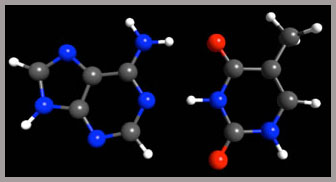
Intermolecular Interaction. We use state-of-the-art computers to perform highly accurate quantum chemical calculations, such as MP2 and CCSD(T), to derive the intermolecular interactions, especially protein-ligand, metal-ligand interactions. We recently implemented a new energy decomposition analysis (EDA) scheme in GAMESS to analyze the interaction in terms of electrostatic, exchange, repulsion, polarization and dispersion, and obtained insights into various molecular systems. Insights into metal-ligand and DNA base pair interactions have been obtained.
Continuum Solvation Model. Solvent effects must be considered in the study of biological molecules. We are extending and improving continuum solvation models for large-scale protein quantum chemical calculations. Recently, we developed a new surface tessellation scheme called FIXPVA for conductorlike continuum models, and obtained rigorously continuous and smooth potential energy surfaces, as well as exact gradients. We have also developed a heterogeneous conductorlike continuum model (Het-CPCM) for quantum chemical studies of protein active sites. These methods are implemented in GAMESS.
For more information, please visit the Li Research Group Homepage.
Selected Publications
(1) Hui Li, Nandun M. Thellamurege, Dejun Si “QuanPol: A QM/MM program for high-level electronic structure methods and polarizable force field”, mss in preparation (24 thousand lines of Fortran code released in GAMESS in Aug 2011)
(2) Dejun Si and Hui Li “Interaction of Aun (n = 1, 2, 4) with sulfur ligands” mss in preparation
(3) Nandun M. Thellamurege and Hui Li “Interaction of Zn, Ca, Mg ions with protein ligands”, mss in preparation