Assistant professor Cliff Stains and his colleagues have led a new study that has shown how the enzyme that sparks bioluminescent fireworks can be used to monitor an indicator of human diseases that include Alzheimer’s, Parkinson’s and Type 2 diabetes. Specifically, the team was able to utilize self-assembling luciferase fragments to quantify the effect of mutations as well as inhibitors on the aggregation of amyloid-beta, a protein whose aggregation is implicated in Alzheimer’s disease pathology.
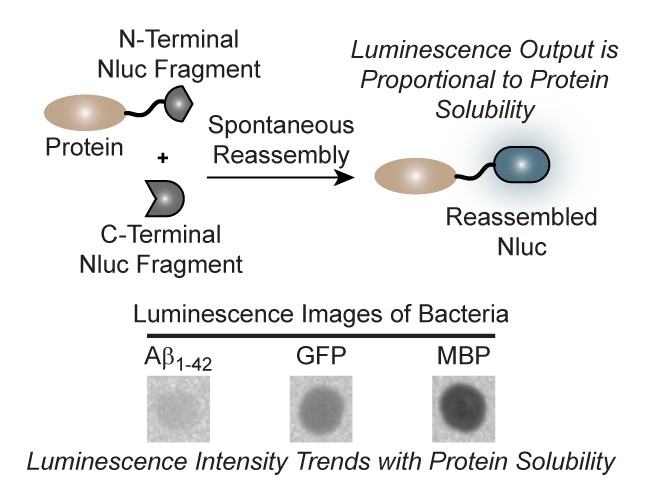
Protein aggregation plays a central role in numerous human disease states including Alzheimer’s (Aβ), Parkinson’s (α-synuclein), and type 2 diabetes (amylin).1 With this in mind, the Stains lab set out to develop a luminescence-based assay for protein aggregation that would be compatible with living cells. As the luminescent reporter, the team chose the recently described nanoluciferase (Nluc) enzyme.2 Utilizing secondary structure prediction algorithms, the research team was able to identify several potential sites of fragmentation within the Nluc enzyme. Interestingly, many of these fragments produced no observable luminescence alone but were capable of spontaneous self-assembly into a functional enzyme when coexpressed in living bacteria (Fig. 1a). Building off of this result, the team characterized purified samples of the optimal fragment pairs, termed N65 and 66C. The results demonstrated that recovered luminescence from this system was directly proportional to the amount of N65 available for reassembly (Fig. 1b). This key result led to the idea that modulating the concentration of the N65 fragment within living cells, by appending an aggregation-prone protein, could lead to a change in cellular luminescence (Fig. 1c). To verify this hypothesis, the researchers fused proteins of known aggregation potential to the N-terminus of N65 and measured the luminescence of bacteria coexpressing 66C. Gratifyingly, a clear trend in luminescence that correlated with the solubility of the protein appended to N65 was observed. Leveraging their new turn-on assay for increased protein solubility the team was able to demonstrate the ability to monitor the effect of mutations as well as small molecules on the aggregation potential of Aβ (Fig. 1d and e). The authors also demonstrated that these assays could be ported into mammalian cells. In the long term, this assay platform could be utilized to screen for inhibitors of protein aggregation as well as identify residues that are essential for aggregation. The group is also pursuing assays for other disease-relevant, aggregation-prone proteins.
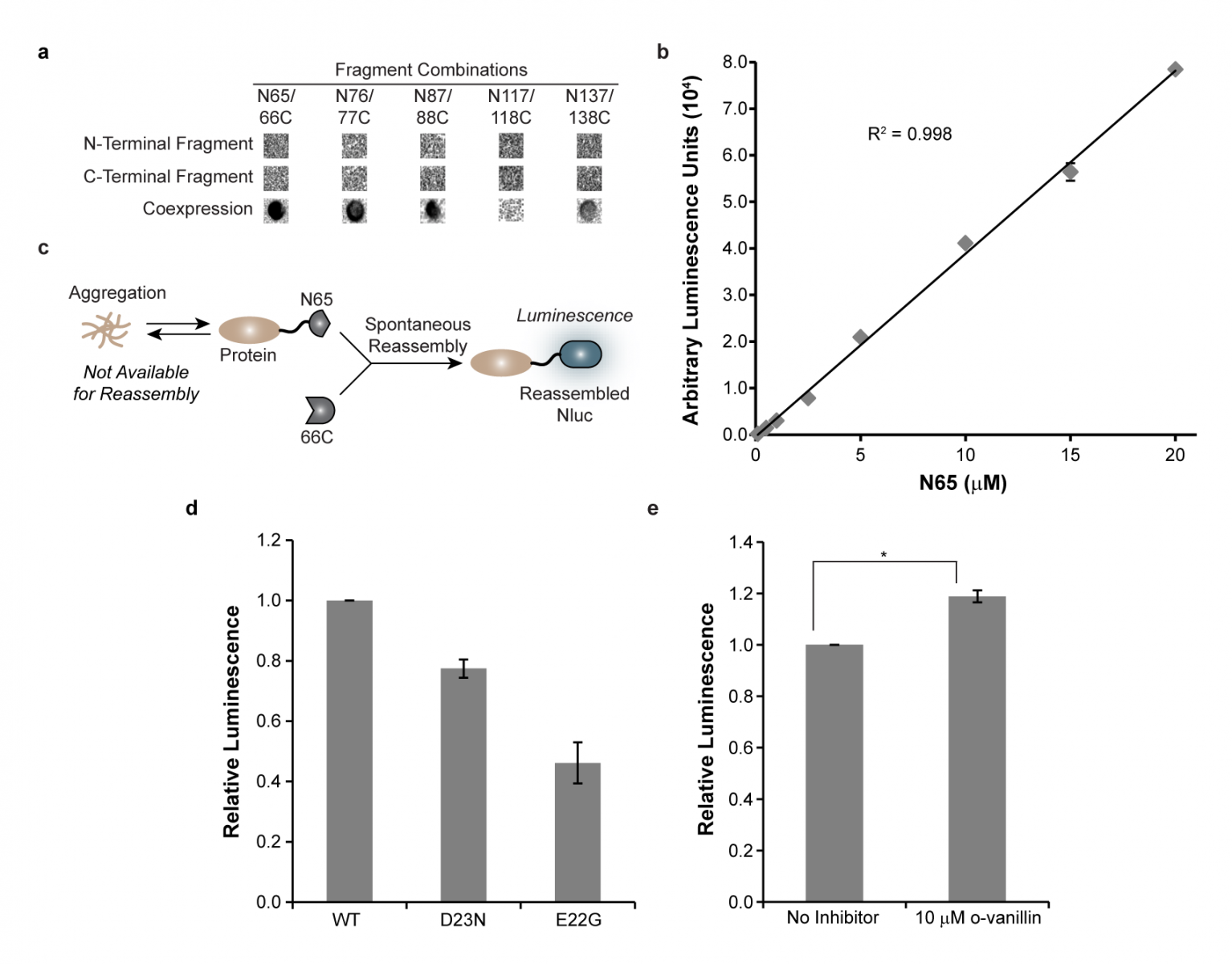
Figure 1. Development of a cellular turn-on luminescence assay for protein solubility. (a) Luminescence of bacterial cells expressing the indicated Nluc fragments. (b) Luminescence signal is proportional to the concentration of N65. (c) Monitoring the solubility of proteins fused to N65 in living cells. Aggregation reduces the concentration of N65 available for reassembly with 66C, leading to a predicable change in luminescence. (d) Luminescence of bacteria expressing wild-type (WT) or the disease-relevant D23N or E22G mutations in Aβ correlate with the known aggregation potential of these proteins. (e) The inhibition of Aβ aggregation by small molecules can be monitored.
The study was published in ACS Chemical Biology and is viewable at https://pubs.acs.org/doi/abs/10.1021/acschembio.5b00758?src=recsys
References:
- Chiti, F. & Dobson, C.M. Protein misfolding, functional amyloid, and human disease. Annu. Rev. Biochem. 75, 333-366 (2006).
- Hall, M.P. et al. Engineered luciferase reporter from a deep sea shrimp utilizing a novel imidazopyrazinone substrate. ACS Chem. Biol. 7, 1848-1857 (2012).



